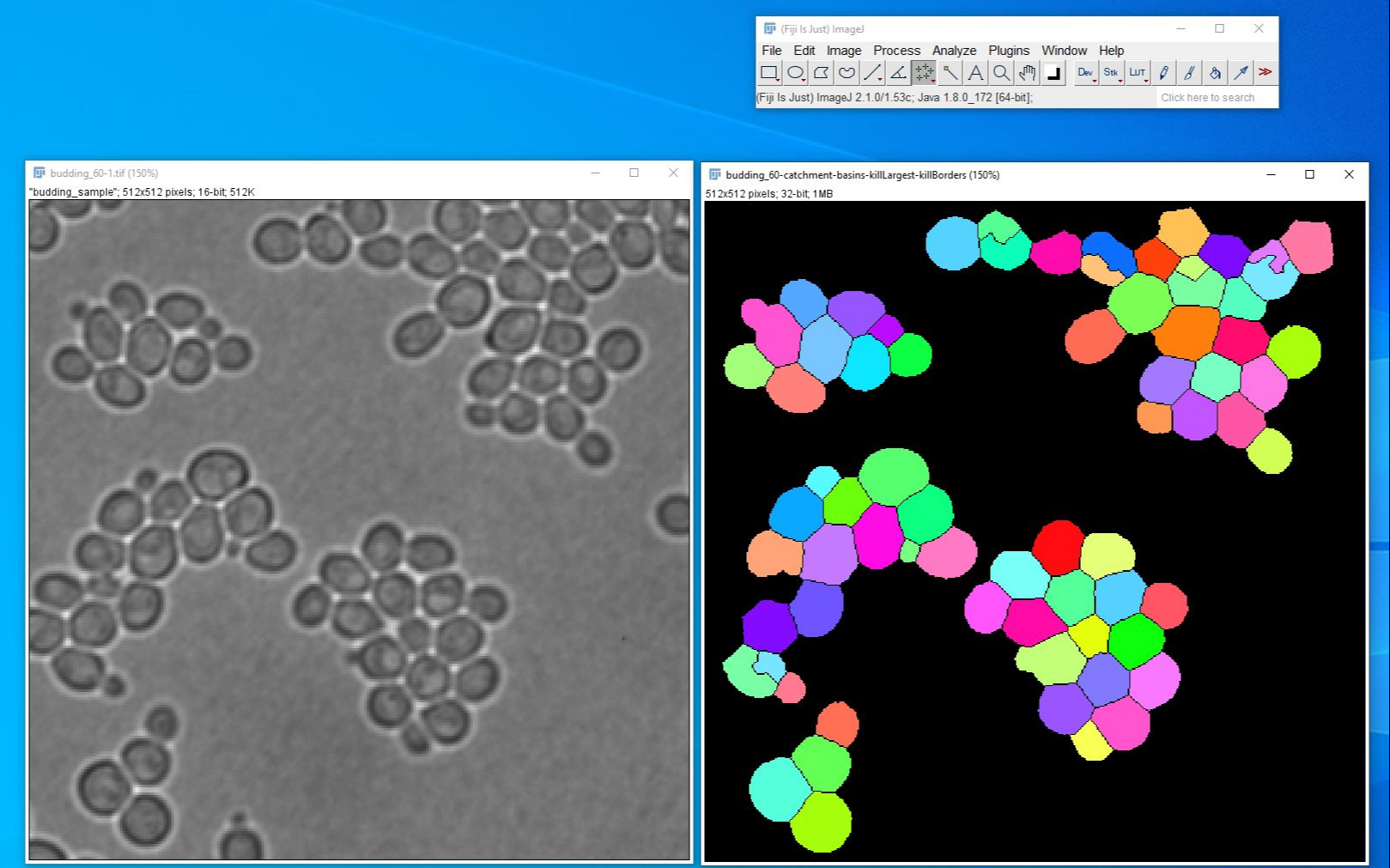
If you have an image of the micrometer available, you draw a line of known length using the line tool, then do the set scale, which will have the line length in pixels already filled in. When all your images were recorded with this scale, tick the Global box as this will apply the Set Scale to all open images. In the Distance in pixels enter 100, in the Known distance enter 645 and the Unit of Length enter µm (um is recognised as µm). Coloc 2 does NOT perform object based colocalization measurements, where objects. It implements and performs the pixel intensity correlation over space methods of Pearson, Manders, Costes, Li and more, for scatterplots, analysis, automatic thresholding and statistical significance testing. To calibrate your image spatially, go to Analyse > Set Scale. Coloc 2 is Fiji’s plugin for colocalization analysis. In the hyperstack window, ensure the number of channels, slices and frames is set correctly (in this example 8 channels, 1 slice, 1. To do this we go to Images -> Hyperstack -> Stack to Hyperstack. We now need to define in FIJI that this is an 8 channel image. When you have the opportunity next time record a micrometer in brightfield using the same microscope settings and verify the size. Note: The images will be added into the stack in the order they were opened in FIJI. The information you could find, was this the pixel size of the camera maybe? Of course it could be the size of a pixel in the image plane, but it seems a bit large. That should easily be verified how big are the monocytes in your image? Having independent depth-cueing for surface (nearest-point) and interior (brightest-point) allows for more visualization possibilities.Monocytes seem to be 15-25 µm in size, so one monocyte then should occupy 2x2 to 4x4 pixels, which is tiny to say the least. For both kinds, depth-cueing is turned off when set to zero (i.e.100% of intensity in back to 100% of intensity in front) and is on when set at 0 < n 100 (i.e.( 100 − n)% of intensity in back to 100% intensity in front). Interior Depth-Cueing works only on brightest-point projections. Surface Depth-Cueing works only on nearest-point projections and the nearest-point component of other projections with opacity turned on. Example: the Fiji/plugins/ActionBar/1pixel. This menu lists all commands related to image processing, including point operations, filters, and arithmetic operations between multiple images 104. 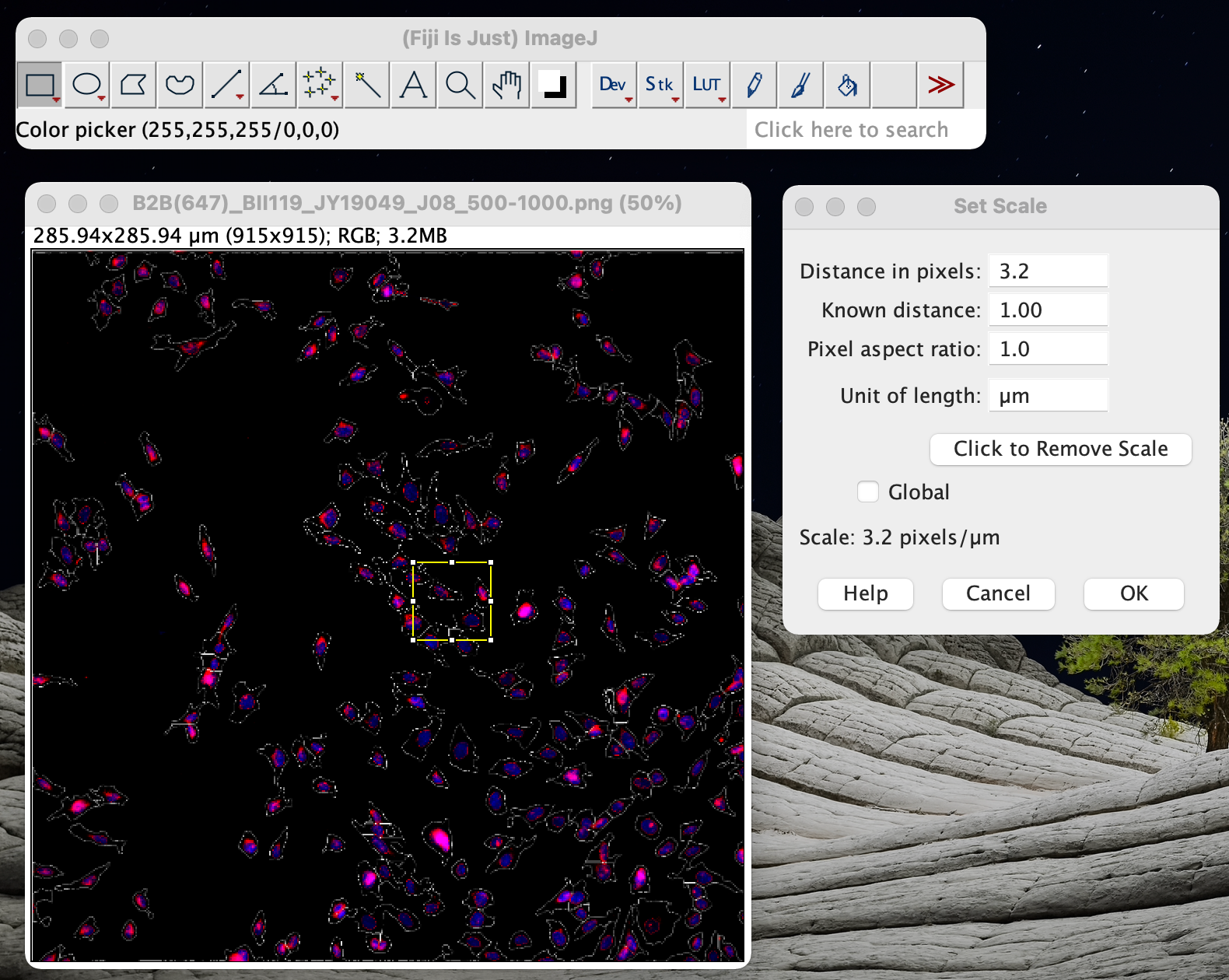
ijm file with an underscore, and refresh menus. The Brightness and Contrast tool may be accessed through Image Adjust Brightness/Contrast or Ctrl / Ctrl on a PC command on a Mac + Shift + C. Two kinds of depth-cueing are available: Surface Depth-Cueing and Interior Depth-Cueing. Works in Fiji : save your custom bars in Fiji/plugins/ActionBar/ as a. This tool may be used to adjust the brightness and contrast of an active image. Cooks call them recipes, biologists protocols, and programmers call them HOWTOs. The trade-off for this increased realism is that data points shown in a depth-cued image no longer possess accurate densitometric values. This section is an introduction to scientific imaging, including both image acquisition and image analysis, with a focus on ImageJ. The depth-cueing parameters determine whether projected points originating near the viewer appear brighter, while points further away are dimmed linearly with distance. For my experiments I have antibody-stained cells across a variety of conditions (wild-type vs. Next select the right stacks for the analysis in Channel1 and Channel2. With the green and red stacks of the colocsample1bRGBBG.tif dataset open and the channels split (see above) choose the menu item Analyze-Colocalization-Colocalization Threshold. Surface/Interior Depth-Cueing Depth cues can contribute to the three-dimensional quality of projection images by giving perspective to projected structures. The Image Stitching package comes with 2 different plugins: Pairwise Stitching: Stitch two 2d-5d images, rectangular ROIs can be used to limit the area to search in. Hi everyone, I’m a new user of ImageJ, and I’m interested in measuring and comparing the fluorescence intensity as part of a time course analysis on embryos at different developmental stages. The Colocalization Threshold plugin performs several functions for you in one go.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
